The exome codes the letters in the encyclopedia. The spaces are important too. -
What is an exome? "Exome" refers to the part of the genome that is coded for making proteins.
Introns are the rest of the genome. This genetic material is NOT coded but serves functions including regulation of expression. We do not know what all the functions are yet.
This article points to a role for introns in causing autism.
Full genome sequencing will eventually be used for detecting these problems.
JR
Spaces near genes may harbor pitfalls for autism
BY JESSICA WRIGHT / 15 FEBRUARY 2016
GENES
DISCUSS THIS ARTICLE
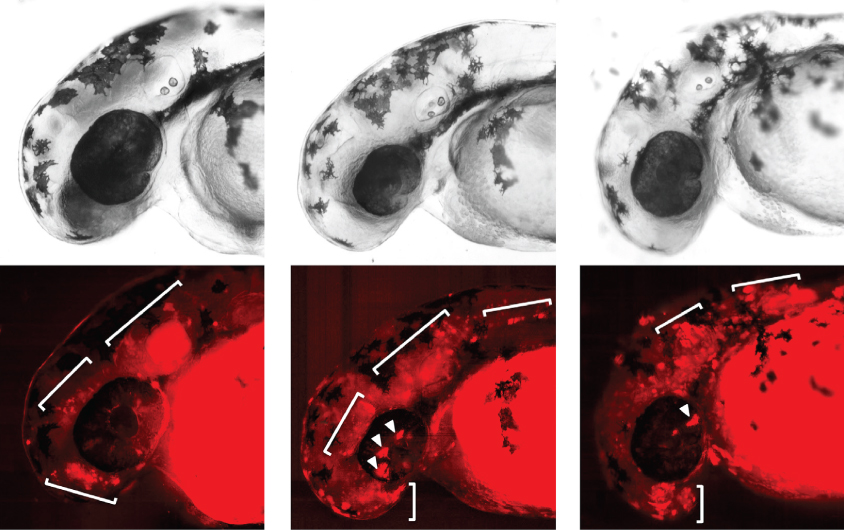
Gene control: Pieces of DNA that are absent in a boy with autism drive expression of a fluorescent protein (red, bottom) in fish brains (top).
Children with autism tend to carry variants in stretches of DNA that flank autism genes. These regions, once dismissed as ‘junk DNA,’ are now known to sometimes control the expression of genes that play key roles in brain development.
The study is small, with just 53 families that each include one person with autism. Still, the findings, published 7 January in the American Journal of Human Genetics, indicate that sequencing whole genomes could reveal the genetic cause of autism in as much as 30 percent of people for whom faster and cheaper sequencing methods come up short1.
“That there [may be] another significant genetic signal in the noncoding regulatory elements is really exciting,” says lead researcher Evan Eichler, professor of genome sciences at the University of Washington in Seattle. “It suggests another class of mutation, which gives us a lot of hope that sequencing whole genomes is going to be worthwhile.”
Eichler and his colleagues scoured sequences of entire genomes — genes and the regions between them — for previously unidentified variants that might be involved in autism. Prior attempts to sequence all the protein-coding regions of the genome, or exomes, had not turned up any autism-linked mutations in these 53 families. The family members also do not carry any large DNA deletions or duplications, called copy number variants (CNVs), known to be involved in autism....
Comparing the sequences of the individuals with autism and those of their unaffected siblings, the researchers found that people with autism are more likely to have genetic variants — either single base-pair changes in the sequence or small CNVs — in swaths of DNA abutting known autism genes. But the researchers only found the variants after they restricted their search to regions of the genome already implicated in autism, and even then the statistical significance is modest.
They found significantly more de novo variants among children with autism than among their siblings between 10 and 100 kilobases on either side of these genes. For example, they found six genetic variants in children with autism within 50 kilobases of the autism genes, but none in their unaffected siblings.
They also looked for CNVs and variants within autism-risk genes that previous sequencing methods might have missed. Whole-genome sequencing can pinpoint small CNVs, whereas traditional methods are usually limited to those larger than 100 kilobases.
Overall, 16 of the 53 people with autism have one or more of these rare variants, meaning they are present in a single person with autism or a single family; nine of the families carry CNVs or mutations that clearly alter genes.
Full article HERE
No comments:
Post a Comment